Number of species of a community
Richness.RdCalculates the number of species from probability vector. The name is that of the estimator (the bias correction) used.
Usage
Richness(NorP, ...)
bcRichness(Ns, Correction = "Best", Alpha = 0.05, JackOver = FALSE, JackMax = 10,
CheckArguments = TRUE)
# S3 method for class 'ProbaVector'
Richness(NorP, ..., CheckArguments = TRUE, Ps = NULL)
# S3 method for class 'AbdVector'
Richness(NorP, Correction = "Best", Alpha = 0.05,
JackOver = FALSE, JackMax = 10,
Level = NULL, PCorrection = "Chao2015", Unveiling = "geom", RCorrection = "Rarefy",
..., CheckArguments = TRUE, Ns = NULL)
# S3 method for class 'integer'
Richness(NorP, Correction = "Best", Alpha = 0.05,
JackOver = FALSE, JackMax = 10,
Level = NULL, PCorrection = "Chao2015", Unveiling = "geom", RCorrection = "Rarefy",
..., CheckArguments = TRUE, Ns = NULL)
# S3 method for class 'numeric'
Richness(NorP, Correction = "Best", Alpha = 0.05,
JackOver = FALSE, JackMax = 10,
Level = NULL, PCorrection = "Chao2015", Unveiling = "geom", RCorrection = "Rarefy",
..., CheckArguments = TRUE, Ps = NULL, Ns = NULL)Arguments
- Ps
A probability vector, summing to 1.
- Ns
A numeric vector containing species abundances.
- NorP
A numeric vector, an integer vector, an abundance vector (
AbdVector) or a probability vector (ProbaVector). Contains either abundances or probabilities.- Correction
A string containing one of the possible corrections:
"None"(no correction),"Jackknife","iChao1", or"Chao1"."Best", the default value, is currently"Jackknife". Ignored by richness interpolation, and by extrapolation ifPCorrectionis not "None".- Alpha
The risk level, 5% by default, used to optimize the jackknife order.
- JackOver
If
TRUE, retain the jackknife order immediately superior to the optimal one, usually resulting in the overestimation of the number of species. Default isFALSE.- JackMax
The highest jackknife order allowed. Default is 10. Allowed values are between 1 and 10.
- Level
The level of interpolation or extrapolation. It may be an a chosen sample size (an integer) or a sample coverage (a number between 0 and 1). Richness extrapolation require its asymptotic estimation depending on the choice of
Correction.- PCorrection
A string containing one of the possible corrections to estimate a probability distribution in
as.ProbaVector:"Chao2015"is the default value. If "None", the asymptotic distribution is not estimated and extrapolation relies only on the asymptotic estimator of richness. Used only for extrapolation.- Unveiling
A string containing one of the possible unveiling methods to estimate the probabilities of the unobserved species in
as.ProbaVector:"geom"(the unobserved species distribution is geometric) is the default value. Used only for extrapolation.- RCorrection
A string containing a correction recognized by
Richnessto evaluate the total number of species inas.ProbaVector."Rarefy"is the default value to estimate the number of species such that the entropy of the asymptotic distribution rarefied to the observed sample size equals the observed entropy of the data. Used only for extrapolation.- ...
Additional arguments. Unused.
- CheckArguments
Logical; if
TRUE, the function arguments are verified. Should be set toFALSEto save time when the arguments have been checked elsewhere.
Details
Bias correction requires the number of individuals. Use bcRichness and choose the Correction.
Chao correction techniques are from Chao (1984) and Chiu et al. (2015). The Jackknife estimator is calculated by a straight adaptation of the code by Ji-Ping Wang (jackknife in CRAN-archived package SPECIES). The optimal order is selected according to Burnham and Overton (1978; 1979). The argument JackOver allows selecting one order over the optimal one.
Many other estimators are available elsewhere, the ones implemented here are necessary for other entropy estimations.
The functions are designed to be used as simply as possible. Richness is a generic method. If its first argument is an abundance vector, an integer vector or a numeric vector which does not sum to 1, the bias corrected function bcRichness is called.
Richness can be estimated at a specified level of interpolation or extrapolation, either a chosen sample size or sample coverage (Chao et al., 2014), rather than its asymptotic value.
Extrapolation relies on the estimation of the asymptotic richness. If PCorrection is "None", then the asymptotic estimation of richness is made using the chosen Correction, else the asymtpotic distribution of the community is derived and its estimated richness adjusted so that the entropy of a sample of this distribution of the size of the actual sample has the entropy of the actual sample.
Value
A named number equal to the estimated number of species.
The name is the Correction, except for "SAC" (Species Accumulation Curve) for interpolation.
References
Burnham, K. P., and Overton,W. S. (1978), Estimation of the Size of a Closed Population When Capture Probabilities Vary Among Animals. Biometrika, 65: 625-633.
Burnham, K. P., and Overton,W. S. (1979), Robust Estimation of Population Size When Capture Probabilities Vary Among Animals. Ecology 60:927-936.
Chao, A. (1984) Nonparametric estimation of the number of classes in a population. Scandinavian Journal of Statistics 11: 265-270.
Chao, A., Gotelli, N. J., Hsieh, T. C., Sander, E. L., Ma, K. H., Colwell, R. K., Ellison, A. M (2014). Rarefaction and extrapolation with Hill numbers: A framework for sampling and estimation in species diversity studies. Ecological Monographs, 84(1): 45-67.
Chiu, C.-H., Wang, Y.-T., Walther, B. A., Chao, A. (2014) An Improved Nonparametric Lower Bound of Species Richness via a Modified Good-Turing Frequency Formula. Biometrics 70(3): 671-682.
Examples
# Load Paracou data (number of trees per species in two 1-ha plot of a tropical forest)
data(Paracou618)
# Ns is the total number of trees per species
Ns <- as.AbdVector(Paracou618.MC$Ns)
# Species probabilities
Ps <- as.ProbaVector(Paracou618.MC$Ns)
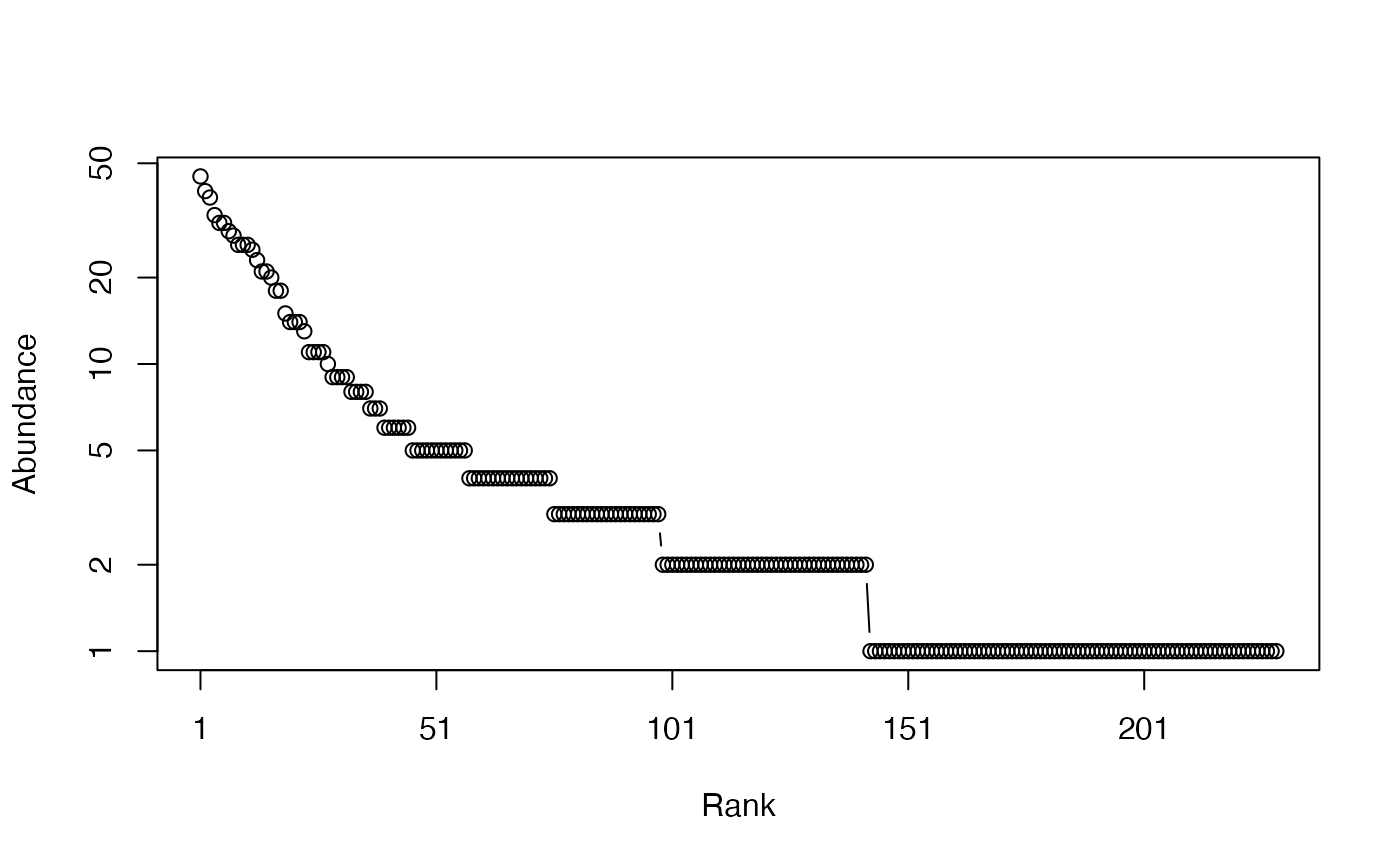
# Whittaker plot
plot(Ns)
 # Number of observed species
Richness(Ps)
#> None
#> 229
# Estimate the actual number of species
bcRichness(Ns, Correction = "Chao1")
#> Chao1
#> 314.9348
bcRichness(Ns, Correction = "iChao1")
#> iChao1
#> 333.7466
bcRichness(Ns, Correction = "Jackknife")
#> Jackknife 2
#> 359
bcRichness(Ns, Correction = "Jackknife", JackOver=TRUE)
#> Jackknife 3
#> 381
# Number of observed species
Richness(Ps)
#> None
#> 229
# Estimate the actual number of species
bcRichness(Ns, Correction = "Chao1")
#> Chao1
#> 314.9348
bcRichness(Ns, Correction = "iChao1")
#> iChao1
#> 333.7466
bcRichness(Ns, Correction = "Jackknife")
#> Jackknife 2
#> 359
bcRichness(Ns, Correction = "Jackknife", JackOver=TRUE)
#> Jackknife 3
#> 381